Southern Blotting is a laboratory technique used to detect a specific DNA sequence within a complex genomic sample. Named after its inventor, Edwin Southern, this method allows scientists to pick out a single “sentence” of genetic information from a library containing millions of words. It is the gold standard for verifying gene insertions, identifying genetic mutations, and was the original foundation for “DNA fingerprinting” in forensic science.
How Southern Blotting Works
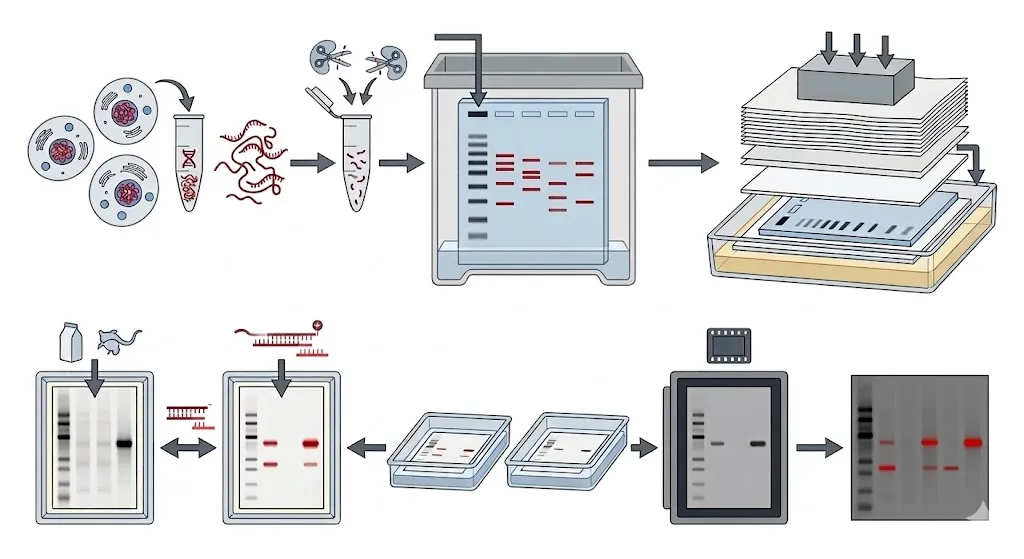
Because a genome is so massive, you can’t just run it on a gel and expect to see individual genes; it would just look like a continuous smear. Southern blotting uses precise cutting and “velcro-like” probes to find the target.
- Digestion
The extracted DNA is treated with restriction enzymes. These molecular scissors cut the DNA only at specific sequences, chopping the long genomic strands into thousands of smaller fragments of varying lengths.
- Electrophoresis
The DNA fragments are separated by size using an agarose gel. Small fragments move quickly through the gel, while large ones stay near the top. To make the DNA “sticky” for the next step, the gel is treated with an alkaline solution to denature the double-stranded DNA into single strands.
- Transfer
The DNA is transferred from the fragile gel onto a sturdy nylon or nitrocellulose membrane. This is typically done through capillary action: a stack of dry paper towels draws SSC (saline-sodium citrate) buffer up through the gel and the membrane. As the liquid moves, it carries the DNA out of the gel and traps it permanently on the membrane’s surface.
- Hybridization and Detection
The membrane is incubated with a labeled DNA probe—a short, single-stranded piece of DNA that is the exact mirror image of the target sequence. The probe “hunts” across the membrane until it finds its match and zips onto it (hybridization). After washing away the extra probe, the membrane is exposed to X-ray film or a digital scanner to reveal the exact location and size of the target gene.